First look of an accepted manuscript (behind paywall), Genome-wide sequence analyses of ethnic populations across Russia, by Zhernakova et al. Genomics (2019).
Interesting excerpts:
There remain ongoing discussions about the origins of the ethnic Russian population. The ancestors of ethnic Russians were among the Slavic tribes that separated from the early Indo-European Group, which included ancestors of modern Slavic, Germanic and Baltic speakers, who appeared in the northeastern part of Europe ca. 1,500 years ago. Slavs were found in the central part of Eastern Europe, where they came in direct contact with (and likely assimilation of) the populations speaking Uralic (Volga-Finnish and Baltic- Finnish), and also Baltic languages [11–13]. In the following centuries, Slavs interacted with the Iranian-Persian, Turkic and Scandinavian peoples, all of which in succession may have contributed to the current pattern of genome diversity across the different parts of Russia. At the end of the Middle Ages and in the early modern period, there occurred a division of the East Slavic unity into Russians, Ukrainians and Belarusians. It was the Russians who drove the colonization movement to the East, although other Slavic, Turkic and Finnish peoples took part in this movement, as the eastward migrations brought them to the Ural Mountains and further into Siberia, the Far East, and Alaska. During that interval, the Russians encountered the Finns, Ugrians, and Samoyeds speakers in the Urals, but also the Turkic, Mongolian and Tungus speakers of Siberia. Finally, in the great expanse between the Altai Mountains on the border with Mongolia, and the Bering Strait, they encountered paleo-Asiatic groups that may be genetically closest to the ancestors of the Native Americans. Today’s complex patchwork of human diversity in Russia has continued to be augmented by modern migrations from the Caucasus, and from Central Asia, as modern economic migrations take shape.
In the current study, we annotated whole genome sequences of individuals currently living on the territory of Russia and identifying themselves as ethnic Russian or as members of a named ethnic minority (Fig. 1). We analyzed genetic variation in three modern populations of Russia (ethnic Russians from Pskov and Novgorod regions and ethnic Yakut from the Sakha Republic), and compared them to the recently released genome sequences collected from 52 indigenous Russian populations. The incidence of function-altering mutations was explored by identifying known variants and novel variants and their allele frequencies relative to variation in adjacent European, East Asian and South Asian populations. Genomic variation was further used to estimate genetic distance and relationships, historic gene flow and barriers to gene flow, the extent of population admixture, historic population contractions, and linkage disequilibrium patterns. Lastly, we present demographic models estimating historic founder events within Russia, and a preliminary HapMap of ethnic Russians from the European part of Russia and Yakuts from eastern Siberia.
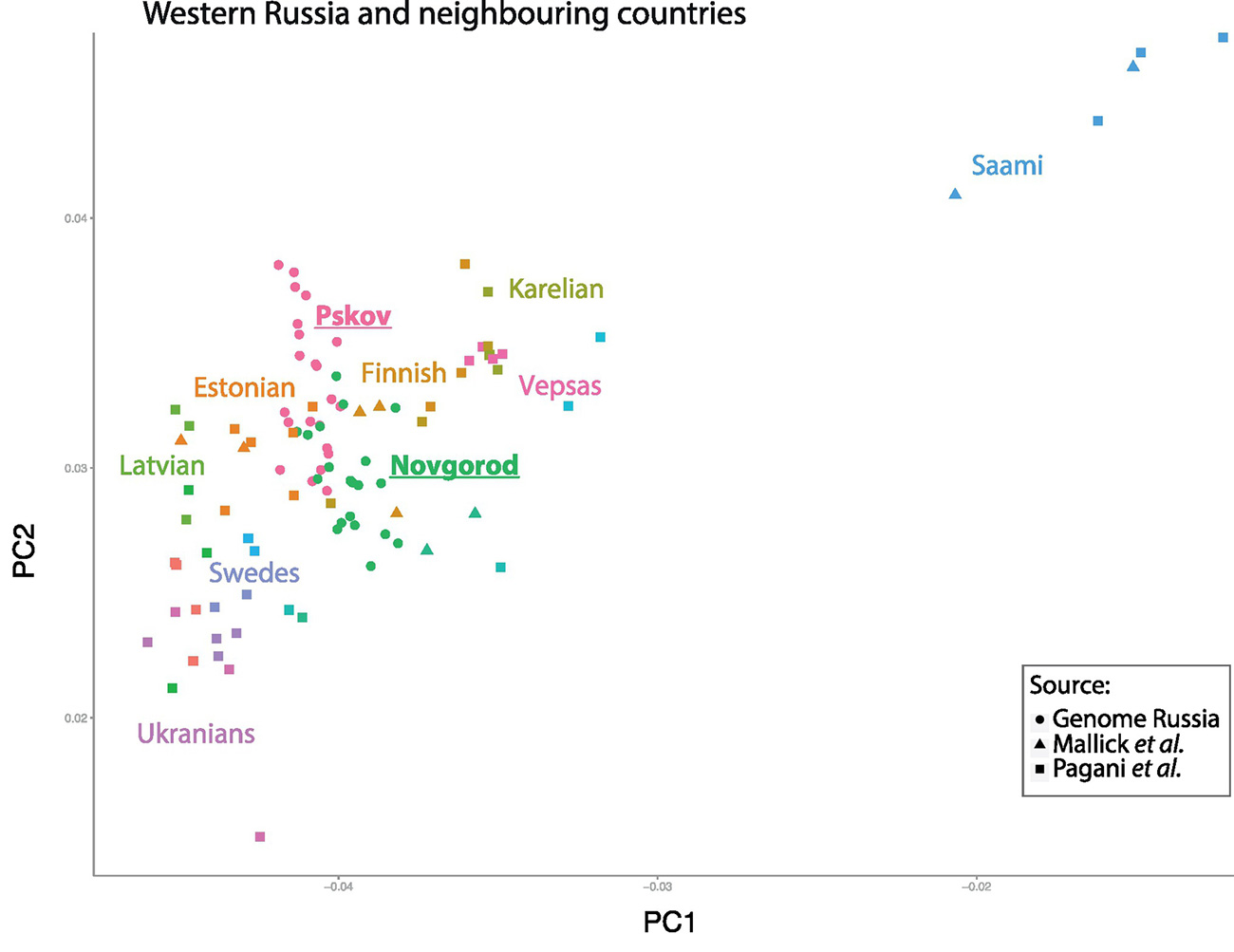
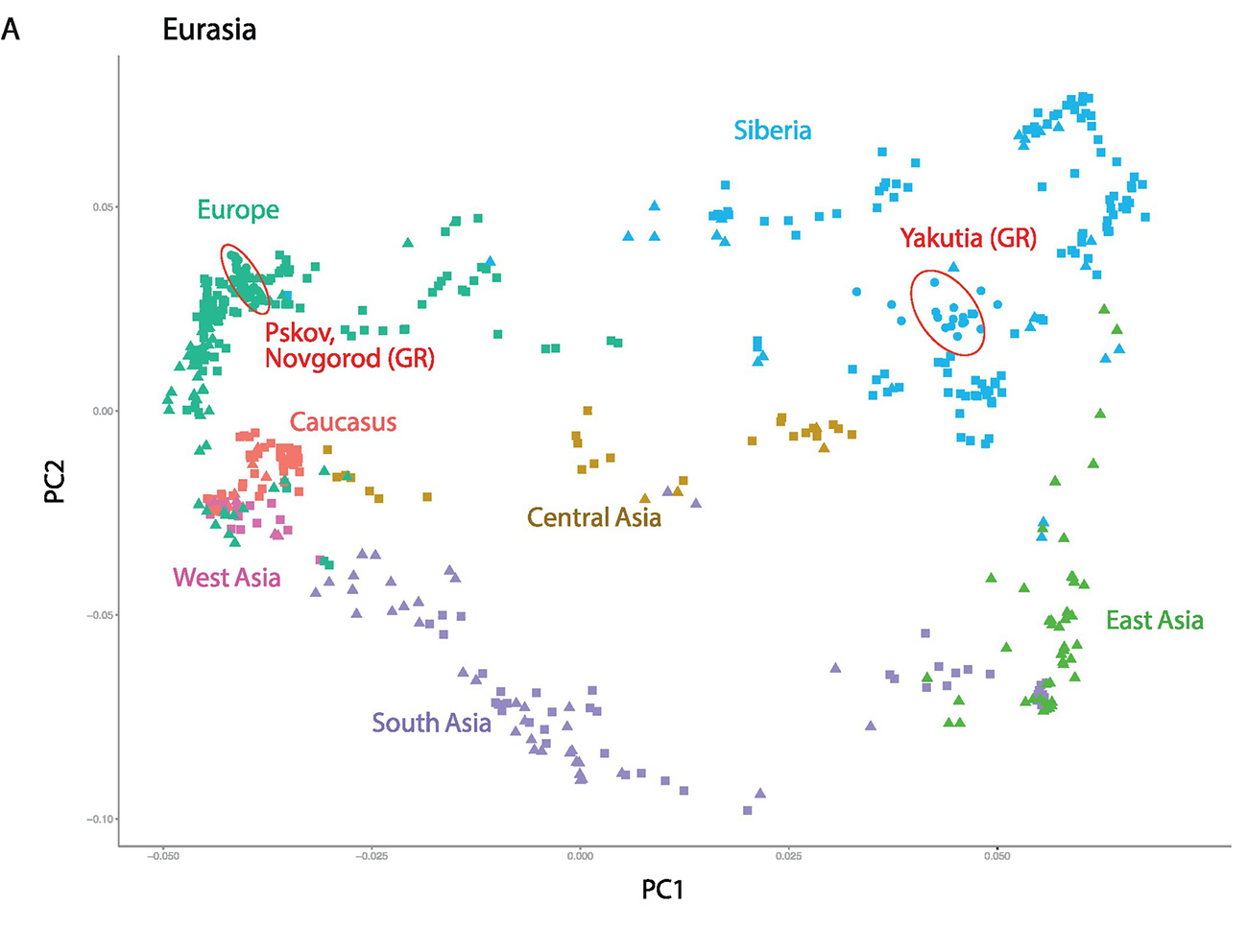
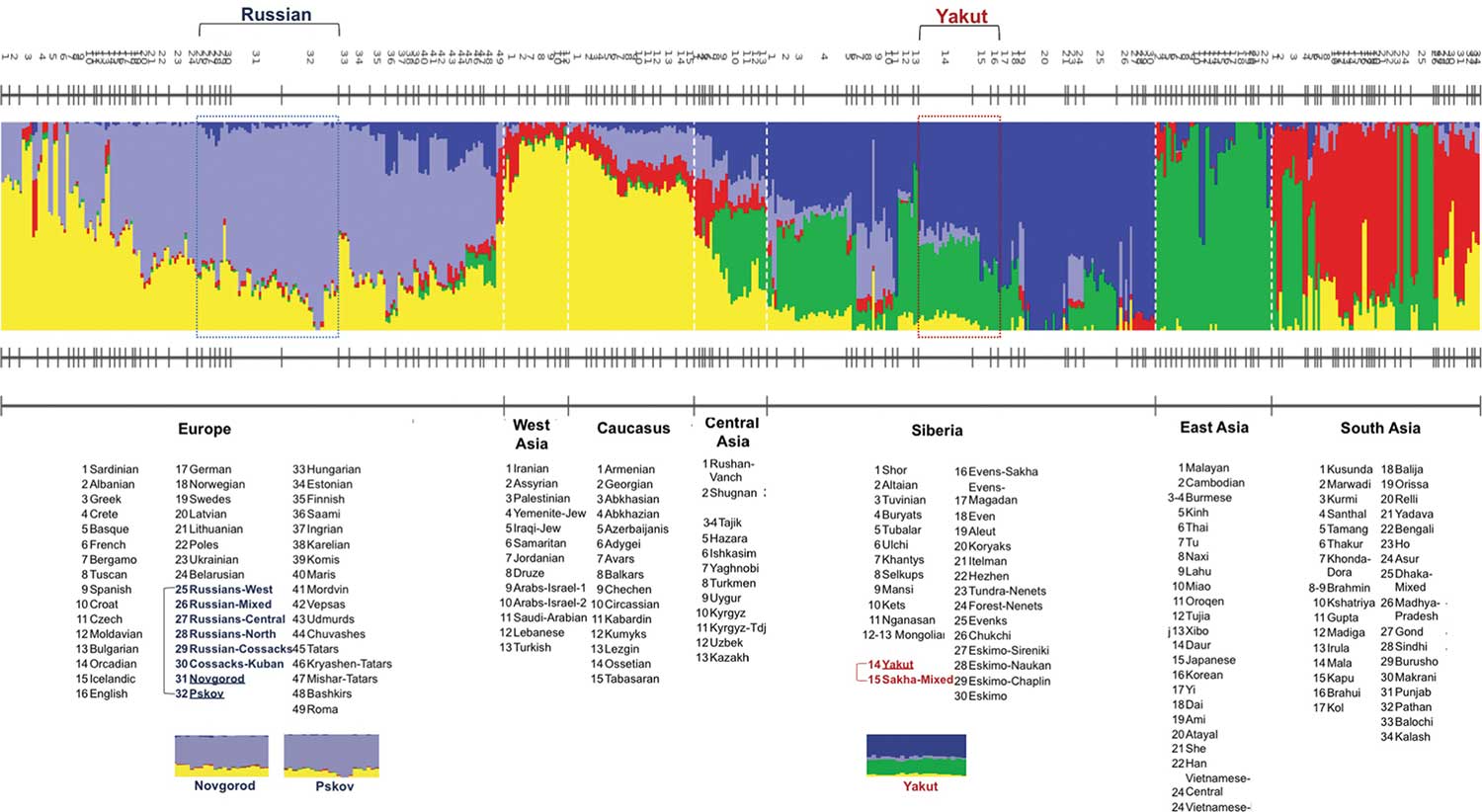
The collection of identified SNPs was used to inspect quantitative distinctions among 264 individuals from across Eurasia (Fig. 1) using Principal Component Analysis (PCA) (Fig. 2). The first and the second eigenvectors of the PCA plot are associated with longitude and latitude, respectively, of the sample locations and accurately separate Eurasian populations according to geographic origin. East European samples cluster near Pskov and Novgorod samples, which fall between northern Russians, Finno-Ugric peoples (Karelian, Finns, Veps etc.), and other Northeastern European peoples (Swedes, Central Russians, Estonian, Latvians, Lithuanians, and Ukrainians) (Fig. 2b). Yakut individuals map into the Siberian sample cluster as expected (Fig. 2a). To obtain an extended view of population relationships, we performed a maximum likelihood-based estimation of ancestry and population structure using ADMIXTURE [46](Fig. 2c). The Novgorod and Pskov populations show similar profiles with their Northeastern European ancestors while the Yakut ethnic group showed mixed ancestry similar to the Buryat and Mongolian groups.
Possible admixture sources of the Genome Russia populations were addressed more formally by calculating F3 statistics, which is an allele frequency-based measure, allowing to test if a target population can be modeled as a mixture of two source populations [48]. Results showed that Yakut individuals are best modeled as an admixture of Evens or Evenks with various European populations (Supplemental Table S4). Pskov and Novgorod showed admixture of European with Siberian or Finno-Ugric populations, with Lithuanian and Latvian populations being the dominant European sources for Pskov samples.
So, Russians expanding in the Middle Ages as acculturated Finno-Volgaic peoples.
Or maybe the true Germano-Slavonic™-speaking area was in north-eastern Europe, until the recent arrival of Finno-Permians with the totally believable Nganasan-Saami horde, whereas Yamna -> Bell Beaker represented Vasconic-Caucasian expanding all over Europe in the Bronze Age. Because steppe ancestry in Fennoscandia and Modern Basques in Iberia.
A really hard choice between equally plausible models.
Related
- Common Slavs from the Lower Danube, expanding with haplogroup E1b-V13?
- Aquitanians and Iberians of haplogroup R1b are exactly like Indo-Iranians and Balto-Slavs of haplogroup R1a
- Magyar tribes brought R1a-Z645, I2a-L621, and N1a-L392(xB197) lineages to the Carpathian Basin
- R1a-Z280 and R1a-Z93 shared by ancient Finno-Ugric populations; N1c-Tat expanded with Micro-Altaic
- Scytho-Siberians of Aldy-Bel and Sagly, of haplogroup R1a-Z93, Q1b-L54, and N
- Mitogenomes from Avar nomadic elite show Inner Asian origin
- The complex origin of Samoyedic-speaking populations
- Corded Ware—Uralic (IV): Hg R1a and N in Finno-Ugric and Samoyedic expansions
- Corded Ware—Uralic (III): “Siberian ancestry” and Ugric-Samoyedic expansions
- Corded Ware—Uralic (II): Finno-Permic and the expansion of N-L392/Siberian ancestry
- The traditional multilingualism of Siberian populations
- Iron Age bottleneck of the Proto-Fennic population in Estonia
- Y-DNA haplogroups of Tuvinian tribes show little effect of the Mongol expansion
- Corded Ware—Uralic (I): Differences and similarities with Yamna
- Haplogroup R1a and CWC ancestry predominate in Fennic, Ugric, and Samoyedic groups